Sponsored content brought to you by
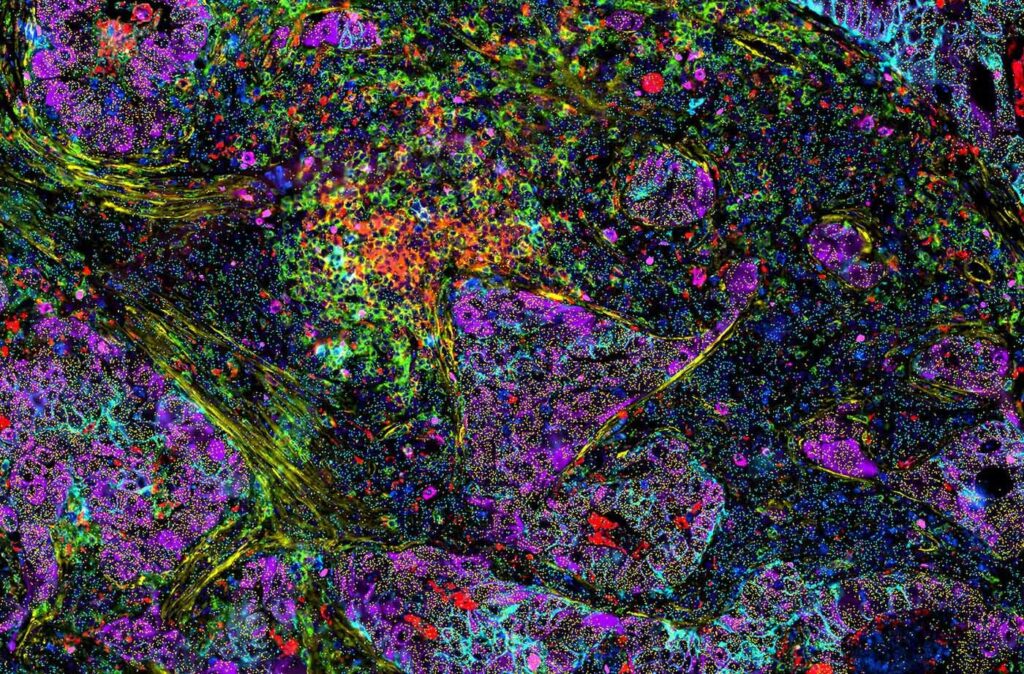
Understanding the composition and architecture of cells within tissues is fundamental to studying tissue function and more broadly, human biology. The complex dynamics of cellular functions and interactions between different cell populations drive normal physiology, while dysregulation of these mechanisms causediseases such as cancer. The recent increase in image-based, spatially resolved technologies enables researchers to profile the tumor microenvironment (TME) by capturing RNA-expression profiles within tissue sections. However, not all RNAs are translated into protein. In some cases, the proteins expressed are exported outside the cell where they affect nearby cells. While the location and identity of RNA and proteins can be visualized with spatial technologies, a significant limitation of these technologies is the lack of ability to resolve protein and RNA information in the same section, as well as conveniently analyze multimodal data sets.
Spatial multiomics made simple

RNAsky® technology is a targeted and amplified spatial gene expression-profiling assay that was designed for the MACSima® Platform. This RNA technology complements the platform’s proven spatial protein detection technology and enables the integration of spatial proteomics and transcriptomics data in the same tissue section. This approach provides in-depth multiomic profiling with single-cell resolution in an automated fashion (Figure 1).
With a broad array of antibodies, RNA detection probes and kits, researchers can develop highly multiplexed assays to thoroughly characterize the TME. In addition, H&E stains can be performed independently on the same section, providing true same-section multiomics.
Analysis of TME populations and spatial dynamics
Spatial biology data sets are often large and complex and can require the assistance of a bioinformatician to analyze the data. However, user-accessible data analysis tools such as MACS® iQ View, can be employed, enabling investigators without an informatics background to analyze data and investigate the impact of clustering using protein, RNA, or both to evaluate TME spatial dynamics. Density and distance map analyses, along with functional characterization of cellular neighborhoods and the cell-to-cell interactions occurring within the TME, can reveal new subpopulations and deepen our understanding of tumor progression in solid tumors.
Practical applications in cancer research
Research presented at recent meetings of the American Association for Cancer Research (AACR) and Society for Immunotherapy of Cancer (SITC) demonstrates the utility of high-plexed same-section spatial multiomics. Scientists interrogated the TME of colorectal cancer (CRC) with a panel of RNA probes and antibodies for deep characterization of the spatial architecture and mechanisms of tumor progression (download the poster). A similar study investigating TME dynamics of CRC revealed the interaction of antigen-presenting cancer-associated fibroblasts (CAFs) and T cells plus it identified a novel immuno-suppressive CAF subpopulation (download the poster).
Multiomic imaging of secondary sarcoma following CAR T-cell therapy for mantle cell lymphoma revealed that the secondary malignancy was not directly associated with the presence of CAR T cells. This study presents a new approach for targeted clinical interventions based on patient biopsies. (See abstract).
Scan the QR code or visit the MACSima Spatial Biology Homepage to learn more.