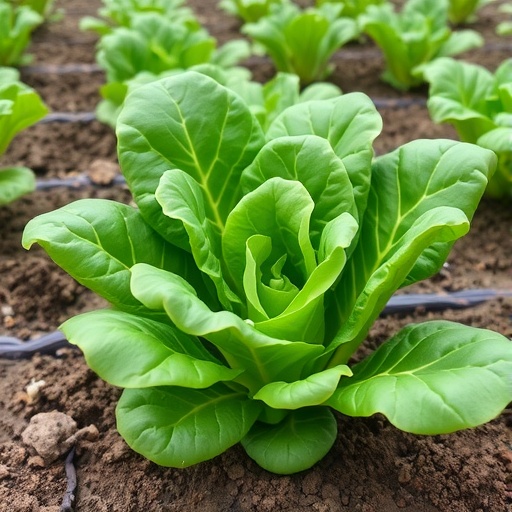
In a pioneering stride toward agricultural sustainability, researchers at the University of Arkansas Division of Agriculture are intensifying efforts to combat the pervasive threat posed by Pythium species to spinach cultivation. This initiative, spearheaded by vegetable breeder Ainong Shi, aims to identify and develop spinach cultivars exhibiting enhanced tolerance to Pythium pathogens, which are notorious for causing root rot and significantly impairing plant vitality. The project is buoyed by a substantial $615,000 grant awarded over three years by the USDA’s National Institute of Food and Agriculture (NIFA), underscoring the critical importance of this research within the agricultural breeding community.
Pythium pathogens, classified as oomycetes—organisms that resemble fungi—pose a particular hazard to spinach by inducing root rot diseases that inhibit nutrient uptake and stunt growth, often culminating in plant death. The challenge is heightened under moist conditions typical of both conventional outdoor fields and controlled environment agriculture systems such as greenhouses and hydroponics. These conditions foster an optimal environment for Pythium proliferation, thereby threatening both yield and quality in various production settings. The Arkansas research team’s focus is to unearth genetic variations that confer partial tolerance, thereby paving the way for cultivars that can withstand these pathogen pressures more robustly.
The complexity of breeding for disease tolerance, especially to a pathogen as insidious as Pythium, stems from the multiplicity of species involved and their interactions with environmental factors. Unlike strategies that seek absolute resistance, this project embraces incremental improvements—enhancing tolerance that reduces disease severity and improves survivability under diverse cultivation conditions. This pragmatic approach acknowledges the polygenic nature of tolerance traits and the dynamic pathogen landscape, advocating for a multifaceted breeding strategy supported by genomic insights.
Central to this endeavor is the development and application of genomic selection methodologies, which enable breeders to predict and select promising genotypes without the lengthy and labor-intensive phenotypic evaluations traditionally required. By harnessing genomic prediction models, Shi’s team accelerates the breeding cycle, identifying candidates with favorable alleles associated with Pythium tolerance. This data-centric approach depends on the rigorous evaluation of hundreds of spinach genotypes—a diverse assemblage drawn from the USDA Agricultural Research Service’s Germplasm Resources Information Network, commercial hybrids and cultivars, as well as Arkansas breeding lines, cumulatively providing a rich genetic tapestry for analysis.
Collaborative integration with industry partners such as 80 Acres Farms, a leader in controlled-environment agriculture, ensures that promising spinach lines not only demonstrate genetic potential but also excel in real-world production contexts. This symbiotic partnership facilitates essential validation of genomic predictions in hydroponic and greenhouse systems, where Pythium challenges are especially acute. Such translational research bridges the gap between laboratory science and commercial agriculture, enhancing the likelihood that newly developed cultivars will meet the needs of growers and consumers alike.
The impetus for this research extends beyond pathogen management; it aligns with broader trends in consumer preferences and market dynamics. Spinach consumption in the United States is on a steady upward trajectory, buoyed by its nutritional profile rich in vitamins A, C, K, folate, iron, calcium, antioxidants, and dietary fiber. This surge is notably driven by the burgeoning fresh-packaged salad industry, which demands consistent quality and supply. Therefore, resilient cultivars that can thrive under both outdoor and indoor production methods are instrumental in supporting this expanding market and meeting year-round consumer demand.
Historically, spinach breeding has prioritized open-field conditions, primarily oriented towards the major growing regions such as California, which accounts for a significant portion of the nation’s production acreage. However, controlled-environment agriculture—encompassing greenhouses, hydroponics, and vertical farms—is gaining traction for its ability to deliver fresh produce near urban populations throughout the year. The project’s focus on improving tolerance to Pythium in these settings acknowledges this shift and addresses the unique pathogen challenges that accompany indoor production systems.
An intriguing dimension of this research is the utilization of a broad genetic pool to capture maximum variation and enhance the prospects of identifying robust tolerance genes. The inclusion of germplasm from diverse sources enriches the breeding base, enabling the mapping of complex traits associated with disease response. This strategy not only benefits spinach breeders but also serves as a model for tackling similar pathogen issues in other vegetable crops, where genetic diversity and genomic tools can be leveraged to expedite cultivar development.
Spinach’s disease management challenges are compounded by the limited availability of cultivars exhibiting high-level resistance to Pythium species. The current reliance on partial tolerance, coupled with agronomic practices aimed at mitigating disease impact, underscores the necessity for genetic innovations. Shi and his team’s genomic research approach represents a paradigm shift, moving beyond traditional breeding to integrate molecular data that informs selection decisions with unprecedented precision and speed.
The Arkansas Agricultural Experiment Station serves as the research nexus for this project, combining expertise from multiple disciplines, including horticulture, plant pathology, and controlled-environment agriculture. The interdisciplinary nature of this team allows for comprehensive analyses—from pathogen biology and host response evaluations to applied breeding and field validation—enhancing the robustness and applicability of research outcomes.
In addressing the pervasive threat of Pythium to spinach production, this research project exemplifies the vital role of public-sector breeding programs in sustaining agricultural productivity against biotic stresses. As consumer demand evolves and production systems diversify, the infusion of genomic technologies into breeding pipelines will enable the development of cultivars that not only meet current agricultural challenges but also anticipate future needs, ensuring food security and nutritional quality.
This groundbreaking initiative embodies a forward-looking strategy that integrates cutting-edge genomic science with practical breeding objectives. It highlights the critical interface between pathogen biology, crop genetics, and production environments, offering a compelling narrative for how modern science can address age-old agricultural challenges. As the project progresses, its findings are expected to catalyze advances in spinach breeding and potentially inspire analogous efforts in other horticultural crops threatened by similar pathogens.
Subject of Research: Development of genomic resources and breeding for Pythium tolerance in spinach.
Article Title: University of Arkansas Researchers Harness Genomic Tools to Breed Pythium-Tolerant Spinach Cultivars.
News Publication Date: Not explicitly stated; inferred to be 2024.
Web References:
– https://www.nifa.usda.gov/about-nifa/announcements/nifa-awards-102m-support-plant-breeding-agricultural-production
– https://portal.nifa.usda.gov/enterprise-search/projects/1034419
– https://aaes.uada.edu/news/pythium-explainer/
– https://www.ers.usda.gov/data-products/vegetables-and-pulses-data/vegetables-and-pulses-yearbook-tables
– https://aaes.uada.edu/
Image Credits: UADA photo.
Keywords: spinach, Pythium, root rot, genomic selection, plant breeding, disease tolerance, controlled-environment agriculture, hydroponics, plant pathology, vegetable breeding, USDA NIFA, Arkansas Agricultural Experiment Station.