In a new study published in Nature titled, “Mapping convergent regulators of melanoma drug resistance by PerturbFate,” researchers from The Rockefeller University have developed a platform called PerturbFate that can systematically map how diverse disease-associated genetic variations reshape cells. By tracking gene regulation in single cells over time, the team identified regulatory nodes common to diverse variations. Using melanoma drug resistance as a proof-of-concept, results showed that these shared points of control offer a path toward combination therapies that can target disease across many genetic causes.
“Once you know that a disease is associated with hundreds of genes, how do you design one therapy to target it?” posed Junyue Cao, PhD, head of the Laboratory of Single-Cell Genomics and Population Dynamics at Rockefeller. “We wondered whether all these different genes may be mediated by some shared downstream signaling that we can discover and target instead.”
Advances in genomic sequencing and genetic screening have allowed researchers to identify hundreds of genetic mutations linked to disease. Yet these genes often span diverse pathways with broad functionalities, from gene regulation to cell signaling, making them difficult to target collectively.
Cao proposed that if these mutations converge on shared downstream programs, the key challenge is not to target each mutation individually, but to identify the common control points known as regulatory nodes.
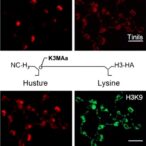
PerturbFate allows researchers to observe how different genetic changes reshape a cell in real time by tracking DNA accessibility, and RNA production and processing. By capturing these changes in the same single cell, the system reveals the networks of genes that control cell behavior and how different genetic variations can have the same effect.
“This technology lets us perturb hundreds to thousands of genes in parallel and then measure the detailed molecular changes in each individual cell,” says Cao. “That allows us to link many different genetic perturbations to their downstream effects and identify regulatory nodes.”
To test the platform, the authors focused on melanoma drug resistance. Using PerturbFate, they selected 143 genes linked to resistance to the common melanoma drug, Vemurafenib. PerturbFate then tracked how deactivating each of these genes reshaped the cell. Cao explains the platform captures gene expression, RNA dynamics and chromatin state, all critical components when identifying upstream regulators that drive these disease states.
After analyzing more than 300,000 cells, the researchers found that diverse genetic perturbations pushed melanoma cells into the same drug-resistant state. Drug resistance dropped significantly when these common control points were targeted, pointing to a promising strategy for combination therapies.
The platform also revealed an important nuance involving the transcriptional coactivator, Mediator Complex. Disrupting different parts of this same complex could trigger drug resistance through routes that ultimately converged on the same survival signal in melanoma cells, called VEGFC. Resistant cells could no longer proliferate after blocking that signal.
The team has made both the experimental and computational tools behind PerturbFate openly available, and plans to extend the approach from cultured cells to living systems. Cao and colleagues are currently applying PerturbFate to conditions, such as aging and Alzheimer’s disease, to uncover shared vulnerabilities that can guide more effective treatments.
“This is just a starting point,” says Cao. “Now that we’ve demonstrated the approach in a simple model, we’re working to extend it into living systems to study even more complex diseases.”